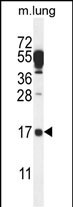
C14orf126 Antibody (C-term)
Affinity Purified Rabbit Polyclonal Antibody (Pab)
- 产品详情
- 实验流程
- 背景知识
Application
| WB, E |
|---|---|
| Primary Accession | Q96FN9 |
| Other Accession | NP_542395.1 |
| Reactivity | Human, Mouse |
| Host | Rabbit |
| Clonality | Polyclonal |
| Isotype | Rabbit IgG |
| Calculated MW | 18660 Da |
| Antigen Region | 136-164 aa |
| Gene ID | 112487 |
|---|---|
| Other Names | Probable D-tyrosyl-tRNA(Tyr) deacylase 2, 31--, D-tyrosyl-tRNA deacylase 2, DTD2, C14orf126 |
| Target/Specificity | This C14orf126 antibody is generated from rabbits immunized with a KLH conjugated synthetic peptide between 136-164 amino acids from the C-terminal region of human C14orf126. |
| Dilution | WB~~1:1000 E~~Use at an assay dependent concentration. |
| Format | Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is purified through a protein A column, followed by peptide affinity purification. |
| Storage | Maintain refrigerated at 2-8°C for up to 2 weeks. For long term storage store at -20°C in small aliquots to prevent freeze-thaw cycles. |
| Precautions | C14orf126 Antibody (C-term) is for research use only and not for use in diagnostic or therapeutic procedures. |
| Name | DTD2 |
|---|---|
| Synonyms | C14orf126 |
| Function | Deacylates mischarged D-aminoacyl-tRNAs (By similarity). Also deacylates mischarged glycyl-tRNA(Ala), protecting cells against glycine mischarging by AlaRS (By similarity). Probably acts by rejecting L-amino acids from its binding site rather than specific recognition of D-amino acids (By similarity). Catalyzes the hydrolysis of D-tyrosyl-tRNA(Tyr), has no activity on correctly charged L-tyrosyl- tRNA(Tyr) (By similarity). By recycling D-aminoacyl-tRNA to D-amino acids and free tRNA molecules, this enzyme counteracts the toxicity associated with the formation of D-aminoacyl-tRNA entities in vivo and helps enforce protein L-homochirality. In contrast to DTD1, deacylates L-Ala mischarged on tRNA(Thr)(G4.U69) by alanine-tRNA ligase AARS (PubMed:29410408). Can deacylate L-Ala due to a relaxed specificity for substrate chirality caused by the trans conformation of the Gly-Pro motif in the active site (PubMed:29410408). Also hydrolyzes correctly charged, achiral, glycyl-tRNA(Gly) in vitro, although in vivo EEF1A1/EF-Tu may protect cognate achiral glycyl-tRNA(Gly) from DTD2- mediated deacetylation (By similarity). |
| Cellular Location | Cytoplasm. |
Research Areas
For Research Use Only. Not For Use In Diagnostic Procedures.
Application Protocols
Provided below are standard protocols that you may find useful for product applications.
BACKGROUND
Hydrolyzes D-tyrosyl-tRNA(Tyr) into D-tyrosine and free tRNA(Tyr). Could be a defense mechanism against a harmful effect of D-tyrosine (Potential).
REFERENCES
Gerhard, D.S., et al. Genome Res. 14 (10B), 2121-2127 (2004) :
终于等到您。ABCEPTA(百远生物)抗体产品。
点击下方“我要评价 ”按钮提交您的反馈信息,您的反馈和评价是我们最宝贵的财富之一,
我们将在1-3个工作日内处理您的反馈信息。
如有疑问,联系:0512-88856768 tech-china@abcepta.com.























 癌症的基本特征包括细胞增殖、血管生成、迁移、凋亡逃避机制和细胞永生等。找到癌症发生过程中这些通路的关键标记物和对应的抗体用于检测至关重要。
癌症的基本特征包括细胞增殖、血管生成、迁移、凋亡逃避机制和细胞永生等。找到癌症发生过程中这些通路的关键标记物和对应的抗体用于检测至关重要。 为您推荐一个泛素化位点预测神器——泛素化分析工具,可以为您的蛋白的泛素化位点作出预测和评分。
为您推荐一个泛素化位点预测神器——泛素化分析工具,可以为您的蛋白的泛素化位点作出预测和评分。 细胞自噬受体图形绘图工具为你的蛋白的细胞受体结合位点作出预测和评分,识别结合到自噬通路中的蛋白是非常重要的,便于让我们理解自噬在正常生理、病理过程中的作用,如发育、细胞分化、神经退化性疾病、压力条件下、感染和癌症。
细胞自噬受体图形绘图工具为你的蛋白的细胞受体结合位点作出预测和评分,识别结合到自噬通路中的蛋白是非常重要的,便于让我们理解自噬在正常生理、病理过程中的作用,如发育、细胞分化、神经退化性疾病、压力条件下、感染和癌症。