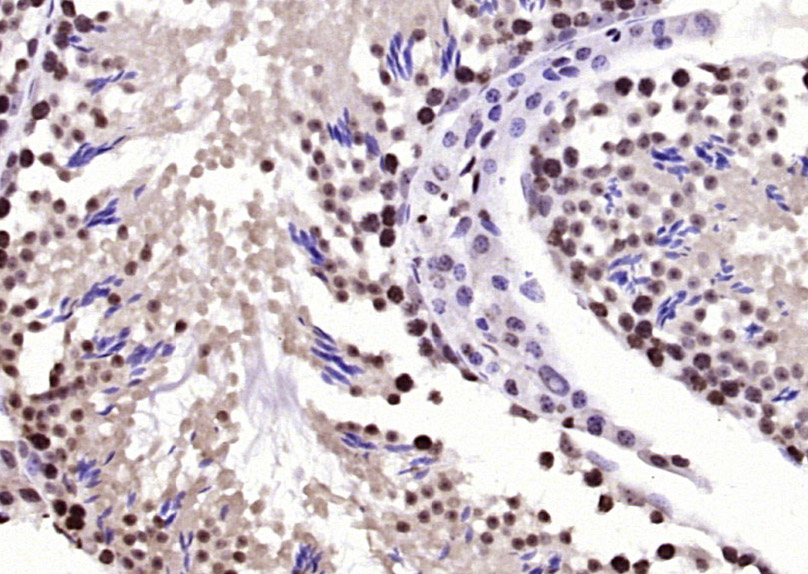
BPTF Rabbit pAb
BPTF Rabbit pAb
- 产品详情
- 实验流程
- 背景知识
Application
| IHC-P, IHC-F, IF |
|---|---|
| Primary Accession | Q12830 |
| Reactivity | Human, Mouse, Rat |
| Predicted | Chicken, Dog, Rabbit, Sheep, Guinea Pig |
| Host | Rabbit |
| Clonality | Polyclonal |
| Calculated MW | 338262 Da |
| Physical State | Liquid |
| Immunogen | KLH conjugated synthetic peptide derived from human BPTF/FALZ |
| Epitope Specificity | 2801-3046/3046 |
| Isotype | IgG |
| Purity | affinity purified by Protein A |
| Buffer | 0.01M TBS (pH7.4) with 1% BSA, 0.02% Proclin300 and 50% Glycerol. |
| SUBCELLULAR LOCATION | Cytoplasm. Nucleus. Note=In brains of Alzheimer disease patients, present in a subset of amyloid-containing plaques. |
| SIMILARITY | Belongs to the PBTF family. Contains 1 bromo domain. Contains 1 DDT domain. Contains 2 PHD-type zinc fingers. |
| SUBUNIT | Interacts with MAZ. Interacts with KEAP1. Part of the nucleosome-remodeling factor (NURF) complex which consists of SMARCA1; BPTF; RBBP4 and RBBP7. Interacts with histone H3K4me3 and to a lesser extent with histone H3-K4Me2. |
| Post-translational modifications | Phosphorylation enhances DNA-binding. Phosphorylated upon DNA damage, probably by ATM or ATR. Highly susceptible to proteolysis. |
| Important Note | This product as supplied is intended for research use only, not for use in human, therapeutic or diagnostic applications. |
| Background Descriptions | This gene was identified by the reactivity of its encoded protein to a monoclonal antibody prepared against brain homogenates from patients with Alzheimer's disease. Analysis of the original protein (fetal Alz-50 reactive clone 1, or FAC1), identified as an 810 aa protein containing a DNA-binding domain and a zinc finger motif, suggested it might play a role in the regulation of transcription. High levels of FAC1 were detected in fetal brain and in patients with neurodegenerative diseases. The protein encoded by this gene is actually much larger than originally thought, and it also contains a C-terminal bromodomain characteristic of proteins that regulate transcription during proliferation. The encoded protein is highly similar to the largest subunit of the Drosophila NURF (nucleosome remodeling factor) complex. In Drosophila, the NURF complex, which catalyzes nucleosome sliding on DNA and interacts with sequence-specific transcription factors, is necessary for the chromatin remodeling required for transcription. Two alternative transcripts encoding different isoforms have been described completely. [provided by RefSeq, Jul 2008] |
| Gene ID | 2186 |
|---|---|
| Other Names | Nucleosome-remodeling factor subunit BPTF, Bromodomain and PHD finger-containing transcription factor, Fetal Alz-50 clone 1 protein, Fetal Alzheimer antigen, BPTF, FAC1, FALZ |
| Target/Specificity | Ubiquitously expressed, with highest levels in testis. Present in kidney, liver and brain. In the brain, highest levels are found in motor cortex (at protein level). |
| Dilution | IHC-P=1:100-500,IHC-F=1:100-500,IF=1:100-500 |
| Storage | Store at -20 °C for one year. Avoid repeated freeze/thaw cycles. When reconstituted in sterile pH 7.4 0.01M PBS or diluent of antibody the antibody is stable for at least two weeks at 2-4 °C. |
| Name | BPTF |
|---|---|
| Synonyms | FAC1, FALZ |
| Function | Regulatory subunit of the ATP-dependent NURF-1 and NURF-5 ISWI chromatin remodeling complexes, which form ordered nucleosome arrays on chromatin and facilitate access to DNA during DNA-templated processes such as DNA replication, transcription, and repair (PubMed:14609955, PubMed:28801535). The NURF-1 ISWI chromatin remodeling complex has a lower ATP hydrolysis rate than the NURF-5 ISWI chromatin remodeling complex (PubMed:28801535). Within the NURF-1 ISWI chromatin-remodeling complex, binds to the promoters of En1 and En2 to positively regulate their expression and promote brain development (PubMed:14609955). Histone-binding protein which binds to H3 tails trimethylated on 'Lys-4' (H3K4me3), which mark transcription start sites of active genes (PubMed:16728976, PubMed:16728978). Binds to histone H3 tails dimethylated on 'Lys-4' (H3K4Me2) to a lesser extent (PubMed:16728976, PubMed:16728978, PubMed:18042461). May also regulate transcription through direct binding to DNA or transcription factors (PubMed:10575013). |
| Cellular Location | Cytoplasm. Nucleus. Note=Localizes to sites of DNA damage (PubMed:25593309). In brains of Alzheimer disease patients, present in a subset of amyloid-containing plaques (PubMed:10727212) |
| Tissue Location | Ubiquitously expressed, with highest levels in testis. Present in kidney, liver and brain. In the brain, highest levels are found in motor cortex (at protein level) |
For Research Use Only. Not For Use In Diagnostic Procedures.
Provided below are standard protocols that you may find useful for product applications.
BACKGROUND
This gene was identified by the reactivity of its encoded protein to a monoclonal antibody prepared against brain homogenates from patients with Alzheimer's disease. Analysis of the original protein (fetal Alz-50 reactive clone 1, or FAC1), identified as an 810 aa protein containing a DNA-binding domain and a zinc finger motif, suggested it might play a role in the regulation of transcription. High levels of FAC1 were detected in fetal brain and in patients with neurodegenerative diseases. The protein encoded by this gene is actually much larger than originally thought, and it also contains a C-terminal bromodomain characteristic of proteins that regulate transcription during proliferation. The encoded protein is highly similar to the largest subunit of the Drosophila NURF (nucleosome remodeling factor) complex. In Drosophila, the NURF complex, which catalyzes nucleosome sliding on DNA and interacts with sequence-specific transcription factors, is necessary for the chromatin remodeling required for transcription. Two alternative transcripts encoding different isoforms have been described completely. [provided by RefSeq, Jul 2008]
终于等到您。ABCEPTA(百远生物)抗体产品。
点击下方“我要评价 ”按钮提交您的反馈信息,您的反馈和评价是我们最宝贵的财富之一,
我们将在1-3个工作日内处理您的反馈信息。
如有疑问,联系:0512-88856768 tech-china@abcepta.com.























 癌症的基本特征包括细胞增殖、血管生成、迁移、凋亡逃避机制和细胞永生等。找到癌症发生过程中这些通路的关键标记物和对应的抗体用于检测至关重要。
癌症的基本特征包括细胞增殖、血管生成、迁移、凋亡逃避机制和细胞永生等。找到癌症发生过程中这些通路的关键标记物和对应的抗体用于检测至关重要。 为您推荐一个泛素化位点预测神器——泛素化分析工具,可以为您的蛋白的泛素化位点作出预测和评分。
为您推荐一个泛素化位点预测神器——泛素化分析工具,可以为您的蛋白的泛素化位点作出预测和评分。 细胞自噬受体图形绘图工具为你的蛋白的细胞受体结合位点作出预测和评分,识别结合到自噬通路中的蛋白是非常重要的,便于让我们理解自噬在正常生理、病理过程中的作用,如发育、细胞分化、神经退化性疾病、压力条件下、感染和癌症。
细胞自噬受体图形绘图工具为你的蛋白的细胞受体结合位点作出预测和评分,识别结合到自噬通路中的蛋白是非常重要的,便于让我们理解自噬在正常生理、病理过程中的作用,如发育、细胞分化、神经退化性疾病、压力条件下、感染和癌症。