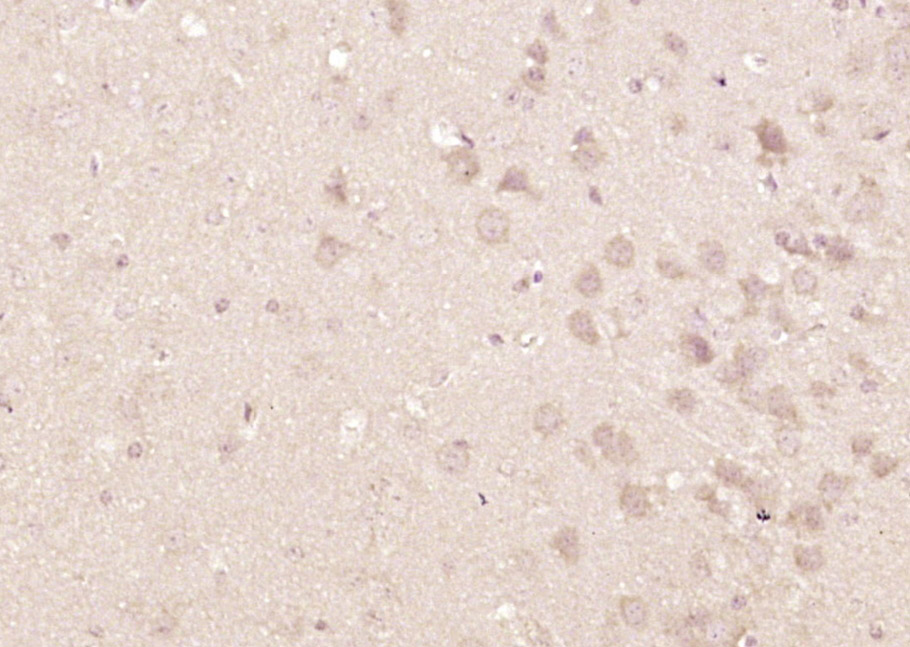
HCF-1 Rabbit pAb
HCF-1 Rabbit pAb
- 产品详情
- 实验流程
- 背景知识
Application
| IHC-P, IHC-F, IF |
|---|---|
| Primary Accession | P51610 |
| Reactivity | Mouse |
| Predicted | Human, Rat, Dog, Pig, Horse |
| Host | Rabbit |
| Clonality | Polyclonal |
| Calculated MW | 208732 Da |
| Physical State | Liquid |
| Immunogen | KLH conjugated synthetic peptide derived from human HCF-1 |
| Epitope Specificity | 201-300/2035 |
| Isotype | IgG |
| Purity | affinity purified by Protein A |
| Buffer | 0.01M TBS (pH7.4) with 1% BSA, 0.02% Proclin300 and 50% Glycerol. |
| SUBCELLULAR LOCATION | Cytoplasm. Nucleus. Note=HCFC1R1 modulates its subcellular localization and overexpression of HCFC1R1 leads to accumulation of HCFC1 in the cytoplasm. Nuclear in general, but uniquely cytoplasmic in trigeminal ganglia, becoming nuclear upon HSV reactivation from the latent state. Non-processed HCFC1 associates with chromatin. |
| SIMILARITY | Contains 5 Kelch repeats. |
| SUBUNIT | Composed predominantly of six polypeptides ranging from110 to 150 kDa and a minor 300 kDa polypeptide. The majority ofN-and C-terminal cleavage products remain tightly, albeitnon-covalently, associated. Interacts with POU2F1, CREB3, ZBTB17,EGR2, E2F4, CREBZF, SP1, GABP2, Sin3 HDAC complex (SIN3A, HDAC1,HDAC2, SDS3), SAP30, SIN3B and FHL2. Component of a MLL1 complex,composed of at least the core components MLL, ASH2L, HCFC1, WDR5and RBBP5, as well as the facultative components C17orf49, CHD8,DPY30, E2F6, HCFC2, HSP70, INO80C, KANSL1, LAS1L, MAX, MCRS1, MEN1,MGA, KAT8, PELP1, PHF20,. PRP31, RING2, RUVBL1, RUVBL2, SENP3,TAF1, TAF4, TAF6, TAF7, TAF9 and TEX10. Component of the MLL5-Lcomplex, composed of at least MLL5, STK38, PPP1CA, PPP1CB, PPP1CC,HCFC1, ACTB and OGT. Component of a THAP1/THAP3-HCFC1-OGT complexthat is required for the regulation of the transcriptional activityof RRM1. Interacts directly with OGT; the interaction, whichrequires the HCFC1 cleavage site domain, glycosylates and promotesthe proteolytic processing of HCFC1, retains OGT in the nucleus andimpacts the expression of herpes simplex virus immediate earlyviral genes. Interacts directly with THAP3 (via its HBM). Interacts(via the Kelch-repeat domain) with THAP1 (via the HBM); theinteraction recruits HCHC1 to the RRM1. Interacts with HCFC1R1 andTHAP11. Associates with the VP16-induced complex; binding to HCFC1activates the viral transcriptional activator VP16 for associationwith POU2F1, to form a multiprotein-DNA complex responsible foractivating transcription of the viral immediate early genes.Component of the SET1 complex, at least composed of the catalyticsubunit (SETD1A or SETD1B), WDR5, WDR82, RBBP5, ASH2L, CXXC1, HCFC1and DPY30. Component of the NSL complex at least composed ofMOF/KAT8, KANSL1, KANSL2, KANSL3, MCRS1, PHF20, OGT1/OGT, WDR5 andHCFC1. |
| Post-translational modifications | Proteolytically cleaved at one or several PPCE--THET siteswithin the HCF repeats. Further cleavage of the primary N- andC-terminal chains results in a 'trimming' and accumulation of thesmaller chains. Cleavage is promoted by O-glycosylation.O-glycosylated. O-glycosylation promotes proteolyticprocessing.Ubiquitinated. Lys-1807 and Lys-1808 are ubiquitinated bothvia 'Lys-48'- and 'Lys-63'-linked polyubiquitin chains. BAP1mediated deubiquitination of 'Lys-48'-linked polyubiquitin chains;deubiquitination by BAP1 does not seem to stabilize the protein. |
| Important Note | This product as supplied is intended for research use only, not for use in human, therapeutic or diagnostic applications. |
| Background Descriptions | This gene is a member of the host cell factor family and encodes a protein with five Kelch repeats, a fibronectin-like motif, and six HCF repeats, each of which contains a highly specific cleavage signal. This nuclear coactivator is proteolytically cleaved at one of the six possible sites, resulting in the creation of an N-terminal chain and the corresponding C-terminal chain. The final form of this protein consists of noncovalently bound N- and C-terminal chains. The protein is involved in control of the cell cycle and transcriptional regulation during herpes simplex virus infection. Alternatively spliced variants which encode different protein isoforms have been described; however, not all variants have been fully characterized. [provided by RefSeq, Jul 2008] |
| Gene ID | 3054 |
|---|---|
| Other Names | Host cell factor 1, HCF, HCF-1, C1 factor, CFF, VCAF, VP16 accessory protein, HCF N-terminal chain 1, HCF N-terminal chain 2, HCF N-terminal chain 3, HCF N-terminal chain 4, HCF N-terminal chain 5, HCF N-terminal chain 6, HCF C-terminal chain 1, HCF C-terminal chain 2, HCF C-terminal chain 3, HCF C-terminal chain 4, HCF C-terminal chain 5, HCF C-terminal chain 6, HCFC1 {ECO:0000303|PubMed:7829097, ECO:0000312|HGNC:HGNC:4839} |
| Target/Specificity | Highly expressed in fetal tissues and the adult kidney. Present in all tissues tested. |
| Dilution | IHC-P=1:100-500,IHC-F=1:100-500,IF=1:100-500 |
| Storage | Store at -20 °C for one year. Avoid repeated freeze/thaw cycles. When reconstituted in sterile pH 7.4 0.01M PBS or diluent of antibody the antibody is stable for at least two weeks at 2-4 °C. |
| Name | HCFC1 {ECO:0000303|PubMed:7829097, ECO:0000312|HGNC:HGNC:4839} |
|---|---|
| Function | Transcriptional coregulator (By similarity). Serves as a scaffold protein, bridging interactions between transcription factors, including THAP11 and ZNF143, and transcriptional coregulators (PubMed:26416877). Involved in control of the cell cycle (PubMed:10629049, PubMed:10779346, PubMed:15190068, PubMed:16624878, PubMed:23629655). Also antagonizes transactivation by ZBTB17 and GABP2; represses ZBTB17 activation of the p15(INK4b) promoter and inhibits its ability to recruit p300 (PubMed:10675337, PubMed:12244100). Coactivator for EGR2 and GABP2 (PubMed:12244100, PubMed:14532282). Tethers the chromatin modifying Set1/Ash2 histone H3 'Lys-4' methyltransferase (H3K4me) and Sin3 histone deacetylase (HDAC) complexes (involved in the activation and repression of transcription, respectively) together (PubMed:12670868). Component of a THAP1/THAP3-HCFC1-OGT complex that is required for the regulation of the transcriptional activity of RRM1 (PubMed:20200153). As part of the NSL complex it may be involved in acetylation of nucleosomal histone H4 on several lysine residues (PubMed:20018852). Recruits KMT2E/MLL5 to E2F1 responsive promoters promoting transcriptional activation and thereby facilitates G1 to S phase transition (PubMed:23629655). Modulates expression of homeobox protein PDX1, perhaps acting in concert with transcription factor E2F1, thereby regulating pancreatic beta-cell growth and glucose-stimulated insulin secretion (By similarity). May negatively modulate transcriptional activity of FOXO3 (By similarity). |
| Cellular Location | Cytoplasm. Nucleus. Note=HCFC1R1 modulates its subcellular localization and overexpression of HCFC1R1 leads to accumulation of HCFC1 in the cytoplasm (PubMed:12235138). Non- processed HCFC1 associates with chromatin. Colocalizes with CREB3 and CANX in the ER. |
| Tissue Location | Highly expressed in fetal tissues and the adult kidney. Present in all tissues tested. |
For Research Use Only. Not For Use In Diagnostic Procedures.
Provided below are standard protocols that you may find useful for product applications.
BACKGROUND
This gene is a member of the host cell factor family and encodes a protein with five Kelch repeats, a fibronectin-like motif, and six HCF repeats, each of which contains a highly specific cleavage signal. This nuclear coactivator is proteolytically cleaved at one of the six possible sites, resulting in the creation of an N-terminal chain and the corresponding C-terminal chain. The final form of this protein consists of noncovalently bound N- and C-terminal chains. The protein is involved in control of the cell cycle and transcriptional regulation during herpes simplex virus infection. Alternatively spliced variants which encode different protein isoforms have been described; however, not all variants have been fully characterized. [provided by RefSeq, Jul 2008]
终于等到您。ABCEPTA(百远生物)抗体产品。
点击下方“我要评价 ”按钮提交您的反馈信息,您的反馈和评价是我们最宝贵的财富之一,
我们将在1-3个工作日内处理您的反馈信息。
如有疑问,联系:0512-88856768 tech-china@abcepta.com.























 癌症的基本特征包括细胞增殖、血管生成、迁移、凋亡逃避机制和细胞永生等。找到癌症发生过程中这些通路的关键标记物和对应的抗体用于检测至关重要。
癌症的基本特征包括细胞增殖、血管生成、迁移、凋亡逃避机制和细胞永生等。找到癌症发生过程中这些通路的关键标记物和对应的抗体用于检测至关重要。 为您推荐一个泛素化位点预测神器——泛素化分析工具,可以为您的蛋白的泛素化位点作出预测和评分。
为您推荐一个泛素化位点预测神器——泛素化分析工具,可以为您的蛋白的泛素化位点作出预测和评分。 细胞自噬受体图形绘图工具为你的蛋白的细胞受体结合位点作出预测和评分,识别结合到自噬通路中的蛋白是非常重要的,便于让我们理解自噬在正常生理、病理过程中的作用,如发育、细胞分化、神经退化性疾病、压力条件下、感染和癌症。
细胞自噬受体图形绘图工具为你的蛋白的细胞受体结合位点作出预测和评分,识别结合到自噬通路中的蛋白是非常重要的,便于让我们理解自噬在正常生理、病理过程中的作用,如发育、细胞分化、神经退化性疾病、压力条件下、感染和癌症。