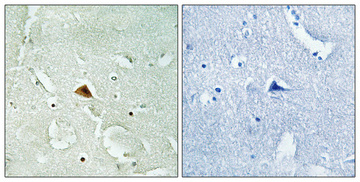
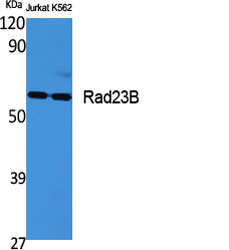
Rad23B Polyclonal Antibody
- 产品详情
- 实验流程
- 背景知识
Application
| WB, IHC-P, IF, ICC, E |
|---|---|
| Primary Accession | P54727 |
| Reactivity | Human, Mouse, Rat |
| Host | Rabbit |
| Clonality | Polyclonal |
| Calculated MW | 43171 Da |
| Gene ID | 5887 |
|---|---|
| Other Names | RAD23B; UV excision repair protein RAD23 homolog B; HR23B; hHR23B; XP-C repair-complementing complex 58 kDa protein; p58 |
| Dilution | WB~~Western Blot: 1/500 - 1/2000. Immunohistochemistry: 1/100 - 1/300. Immunofluorescence: 1/200 - 1/1000. ELISA: 1/20000. Not yet tested in other applications. IHC-P~~Western Blot: 1/500 - 1/2000. Immunohistochemistry: 1/100 - 1/300. Immunofluorescence: 1/200 - 1/1000. ELISA: 1/20000. Not yet tested in other applications. IF~~1:50~200 ICC~~N/A E~~N/A |
| Format | Liquid in PBS containing 50% glycerol, 0.5% BSA and 0.09% (W/V) sodium azide. |
| Storage Conditions | -20℃ |
| Name | RAD23B (HGNC:9813) |
|---|---|
| Function | Multifunctional protein that participates in histone H4K20 demethylation, DNA repair, ubiquitin-dependent protein degradation and transcriptional regulation (PubMed:10488153, PubMed:32209475, PubMed:9372924). Specifically demethylates mono-, di- and trimethylated 'Lys-20' of histone H4 (H4K20me1, H4K20me2, H4K20me3, respectively) into unmethylated forms. Activates the transcription of coding genes by demethylating H4K20me1 and the transcription of repetitive elements by demethylating H4K20me3 (PubMed:32209475). Multiubiquitin chain receptor involved in modulation of proteasomal degradation. Binds to polyubiquitin chains. Proposed to be capable to bind simultaneously to the 26S proteasome and to polyubiquitinated substrates and to deliver ubiquitinated proteins to the proteasome (PubMed:10488153, PubMed:19435460). May play a role in endoplasmic reticulum-associated degradation (ERAD) of misfolded glycoproteins by association with PNGase and delivering deglycosylated proteins to the proteasome (PubMed:15358861). Involved in global genome nucleotide excision repair (GG-NER) by acting as component of the XPC complex, a nucleotide- excision repair complex that is involved in damage sensing during global genome nucleotide excision repair. In vitro, the XPC:RAD23B dimer is sufficient to initiate NER; it preferentially binds to cisplatin and UV-damaged double-stranded DNA. Recognizes a wide spectrum of damaged DNA characterized by distortions of the DNA helix including single-stranded loops, mismatched bubbles or single-stranded overhangs. Cooperatively with CETN2 appears to stabilize XPC (PubMed:10873465, PubMed:12815074, PubMed:9372924). |
| Cellular Location | Nucleus. Cytoplasm Note=The intracellular distribution is cell cycle dependent. Localized to the nucleus and the cytoplasm during G1 phase. Nuclear levels decrease during S-phase; upon entering mitosis, relocalizes in the cytoplasm without association with chromatin |
| Tissue Location | [Isoform 2]: Highly expressed in the testis and in ejaculated spermatozoa. |
For Research Use Only. Not For Use In Diagnostic Procedures.
Provided below are standard protocols that you may find useful for product applications.
BACKGROUND
Multiubiquitin chain receptor involved in modulation of proteasomal degradation. Binds to polyubiquitin chains. Proposed to be capable to bind simultaneously to the 26S proteasome and to polyubiquitinated substrates and to deliver ubiquitinated proteins to the proteasome. May play a role in endoplasmic reticulum- associated degradation (ERAD) of misfolded glycoproteins by association with PNGase and delivering deglycosylated proteins to the proteasome. The XPC complex is proposed to represent the first factor bound at the sites of DNA damage and together with other core recognition factors, XPA, RPA and the TFIIH complex, is part of the pre-incision (or initial recognition) complex. The XPC complex recognizes a wide spectrum of damaged DNA characterized by distortions of the DNA helix such as single-stranded loops, mismatched bubbles or single-stranded overhangs. The orientation of XPC complex binding appears to be crucial for inducing a productive NER. XPC complex is proposed to recognize and to interact with unpaired bases on the undamaged DNA strand which is followed by recruitment of the TFIIH complex and subsequent scanning for lesions in the opposite strand in a 5'-to-3' direction by the NER machinery. Cyclobutane pyrimidine dimers (CPDs) which are formed upon UV-induced DNA damage esacpe detection by the XPC complex due to a low degree of structural perurbation. Instead they are detected by the UV-DDB complex which in turn recruits and cooperates with the XPC complex in the respective DNA repair. In vitro, the XPC:RAD23B dimer is sufficient to initiate NER; it preferentially binds to cisplatin and UV-damaged double-stranded DNA and also binds to a variety of chemically and structurally diverse DNA adducts. XPC:RAD23B contacts DNA both 5' and 3' of a cisplatin lesion with a preference for the 5' side. XPC:RAD23B induces a bend in DNA upon binding. XPC:RAD23B stimulates the activity of DNA glycosylases TDG and SMUG1.
终于等到您。ABCEPTA(百远生物)抗体产品。
点击下方“我要评价 ”按钮提交您的反馈信息,您的反馈和评价是我们最宝贵的财富之一,
我们将在1-3个工作日内处理您的反馈信息。
如有疑问,联系:0512-88856768 tech-china@abcepta.com.























 癌症的基本特征包括细胞增殖、血管生成、迁移、凋亡逃避机制和细胞永生等。找到癌症发生过程中这些通路的关键标记物和对应的抗体用于检测至关重要。
癌症的基本特征包括细胞增殖、血管生成、迁移、凋亡逃避机制和细胞永生等。找到癌症发生过程中这些通路的关键标记物和对应的抗体用于检测至关重要。 为您推荐一个泛素化位点预测神器——泛素化分析工具,可以为您的蛋白的泛素化位点作出预测和评分。
为您推荐一个泛素化位点预测神器——泛素化分析工具,可以为您的蛋白的泛素化位点作出预测和评分。 细胞自噬受体图形绘图工具为你的蛋白的细胞受体结合位点作出预测和评分,识别结合到自噬通路中的蛋白是非常重要的,便于让我们理解自噬在正常生理、病理过程中的作用,如发育、细胞分化、神经退化性疾病、压力条件下、感染和癌症。
细胞自噬受体图形绘图工具为你的蛋白的细胞受体结合位点作出预测和评分,识别结合到自噬通路中的蛋白是非常重要的,便于让我们理解自噬在正常生理、病理过程中的作用,如发育、细胞分化、神经退化性疾病、压力条件下、感染和癌症。
.jpg)